Projects

A01
For the development of novel adjuvant therapies to prevent early metastatic colony formation and to eradicate candidate metastasis founders, it is essential to understand the genetic, epigenetic and functional armament of disseminated cancer cells (DCCs). In project A01, we will address this by a three-step approach. Firstly, we will establish cell lineage trees of longitudinally collected samples from patients progressing from M0-stage disease to M1-stage disease. We will identify the DCC(s) located closest to the root of the metastatic branch of the cell lineage tree and perform detailed genetic characterisation. Secondly, we will mine the transcriptomes and genomes of DCCs to identify molecular mechanisms associated with early metastatic colonisation. Thirdly, we will develop cellular models representative of metastasis founders for functional evaluation and therapeutic targeting.
A02
Tumor progression and metastatic colonization are dynamic processes. Some events happen early in history, others late. Some events can set the stage for colonization of a distant organ, while others such as driver mutations can change the course of a tumor’s history entirely. In the public domain we have an atlas of these events, here we want to compliment it by a history book. We will use our algorithm MHN to annotate mutations by their timing (early/late), by a cancer driver score, and by possible associations with metastasis. To include immunological and clinical events we collaborate with the ongoing PRAEGNANT breast cancer study which generates sequences of up to 850 pairs of primary tumor and metastasis genomes together with immunological characterizations and clinical follow up.
A03
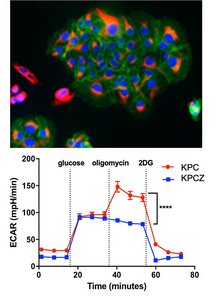
Metastatic colonisation requires a dynamic adaption of cancer cells to the permanently changing conditions of the tumour environment. This is enabled by the activation of the embryonic epithelial to mesenchymal (EMT) program, which provides the necessary plasticity to cancer cells. We and others have demonstrated that his plasticity also includes a permanent adaptation of metabolic processes. In project A03 we will address how EMT-activation is coupled to metabolic changes in metastatic colonisation, with the aim to identify molecular bottlenecks as novel therapeutic targets. We will identify and characterize how EMT factors regulate metabolic processes (e.g. a switch between central energy consumption pathways), explore ways of targeted interference and validate the findings in human cancer.
A04
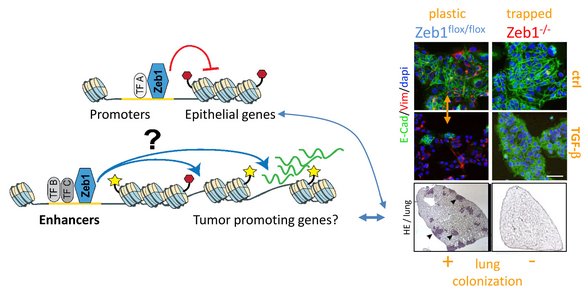
Tumour cell plasticity is a crucial feature for metastatic colonisation. The EMT-transcription factor Zeb1 is mediating this plasticity, and driving colonisation and metastasis, a process accompanied by substantial enhancer reprogramming. Zeb1 knock-out cell lines from a genetic mouse model of pancreatic cancer (KPC) are trapped in an epithelial phenotype and exhibit very low lung colonisation capacity compared to plastic Zeb1 wt lines. Since Zeb1 functions as a transcriptional co-activator mainly in putative enhancer regions, we aim to characterise the enhancer landscape of the above-mentioned cell lines to identify Zeb1 dependent enhancers and their target genes and test their relevance for colonisation and metastasis. We will validate our findings in human cell lines and organoids, thus advancing our understanding about molecular mechanisms of metastasis, which will provide the prospect of novel prognostic tools and putative targets for therapeutic strategies.
A05
We have recently found that the chromatin-remodeling factor Baz2a is highly upregulated in migratory breast cancer cells that display stemness properties. Furthermore, Baz2a is upregulated in small micrometastases, and downregulated in highly proliferative advanced primary tumors and metastases. Baz2a is a subunit of the chromatin remodeling complex, NoRC, that plays an important role as the epigenetic regulator of gene expression. Baz2a binds to a non-coding RNA (pRNA) that regulates the function and genomic targeting of the complex. We hypothesize that Baz2a levels and pRNA processing play important roles in the regulation of cancer cell plasticity by changing gene expression programs and phenotypes of cancer cells, enabling metastatic colonization. We aim to address the molecular role of Baz2a containing remodeling complexes in the process of metastasis, addressing its role in inducing stemness and survival of breast cancer cells after dissemination, and its impact on colonization at the distant site.
A06
The development of metastases from disseminated cancer cells requires adaptation to potentially hostile environments. One factor creating such a hostile environment is chemotherapy, e.g. adjuvant chemotherapy after removal of a primary malignancy. Adaptation of early disseminated cancer cells and formation of metastatic colonies resulting in recurrent disease is a main reason for cancer mortality. In this setting, low-dose metronomic therapy (LDM), which is the daily, long-term low-dose administration of chemotherapeutic drugs, has been successful in patients refractory to standard chemotherapy. Non-coding RNAs play a crucial role during colonization of disseminated cancer cells and the development of cancer metastasis under LDM. We found several lncRNAs that encode for micropeptides, most of them with unknown functions. Project A06 will assess how non-coding RNAs and micropeptides derived from lncRNAs affect metastatic colony formation during LDM. We hypothesize that the highly complex and plastic ‘non-coding’ transcriptome is an important regulator of metastatic colonization and response to or escape from chemotherapy.
A07
To support the discovery of clinically relevant treatments against metastatic progression, it is the goal of this project to develop complex 3D in vitro models and tailored cell-based assays. These will be used to emulate and study the fundamental processes during early metastasis for later use in drug screening campaigns. The project comprises: (i) Development of patient-derived 3D tumor organoid models including their adaptation to high-throughput and high-content screening requirements; (ii) Establishment of physiologically relevant, homotypic or heterotypic iPS-based in vitro models that represent metastatic niches most relevant to breast, pancreas, melanoma and colorectal cancers; (iii) Extending and combining the range of available readout technologies to monitor the 3D nichemodels during experimental encounter of tumor cells and during drug exposure. The experimental readouts include target-based endpoint assays and phenotypic, imaged-based or integral, label-free approaches developed into a modular platform.
B01
Tumour-infiltrating cells of the myeloid compartment are crucial determinants of tumour progression towards metastasis. Particularly tumor associated macrophages (TAMs) where described to exert both tumour/metastasis promoting and suppressing functions. The underlying molecular basis is not fully understood. Our preliminary data indicate that the EMT transcription factor Zeb1 defines a colonization-suppressive subpopulation of TAMs. The aim in project B01 is understand, how Zeb1 determines the generation of TAM-subpopulations, to isolate and characterize the Zeb1+TAM subpopulation in a mouse model for metastatic colonisation, and to validate the findings in human cancer.
B02
Tumor-immune cell interactions can modulate tumor establishment and metastasis. Depending on the cytokine environment macrophages and other myeloid cells can fuel inflammation or trigger resolution of inflammation resulting in suppression of (tumor)immunity and tissue repair. In line with this notion, tumor associated macrophages (TAM) can support or limit tumor growth and metastasis depending on the organ environment. The project aims at (1) identifying organ specific roles of bone marrow derived versus tissue resident macrophages in modulating tumor growth and early metastatic colonisation, (2) at characterizing and alter the phenotype of TAM subpopulations present in tumors growing in different organ environments, and (3) at studying the role of myeloid effector cells in supporting or limiting human primary melanoma and early metastatic melanoma colonisation and growth in humanized mice in vivo.
B03
We have previously identified a conserved “danger containing” Trem2+ macrophage population, which emerged from data of single cell sequencing and preclinical models across various states of inflammation and cancer. These Trem2+ macrophages display a unique immune-regulatory gene expression profile and seem to be able to contain damage and prevent inflammation and local immunity by physical segregation. In project B03, we aim to study the role and origin of these cells during metastatic organ colonization using state of the art imaging, fate-mapping, spatial transcriptomics and single cell RNA sequencing to validate their potential as targets for therapeutic intervention during metastatic colonization.
B04
Regulatory T cells (Treg cells) play an important role in initiating a pre-metastatic niche, promoting metastasis and metastatic tumor growth. They can be induced by a novel cell subset of bone marrow-resident Aire+ tolerogenic antigen presenting cells (BMACs) that express tissue-restricted self-antigens. Within this project we will study a role of BMACs as key regulators of local and systemic immune tolerance towards metastatic cancer cells and establishment of early tumor metastasis. We will use unique transgenic reporter mice to i) unravel the role of BMAC-induced Treg cells in tumor metastasis and BM-colonization, ii) determine to what degree BMACs contribute to the Treg repertoire, iii) identify mechanisms of Treg cell induction by BMACs and iv) explore mechanisms underlying Aire expression in BMACs. The results will open avenues to inhibit metastasis promoting Treg cells.
B05
The formation of metastases in malignant melanoma is associated with a very bad prognosis for melanoma patients. Dendritic cells (DCs) exert an important role in the induction and guidance of innate and adaptive immune responses. Although the role of DCs is started to be investigated in solid primary tumours, their precise role during the metastatic colonization and the unscathed growth of melanoma metastases in a new tissue microenvironment is largely unknown. In project B05 we want to understand the immune control of metastatic colonization by DCs. We will characterize different DC subtypes after isolating infiltrating DCs from metastases, compare them to DCs from primary tumours and corresponding normal tissues, define their specific immune-modulatory effects on T cell subpopulations in in vitro and in vivo models and explore DC subpopulations for therapeutic approaches.
B06
Compelling evidence has accumulated that IL-33, a member of the IL-1 family of cytokines, and its corresponding receptor ST2 are important in cancer progression and metastasis. However, the cellular source of IL-33 as well as the cellular and molecular signaling pathways induced in ST2-positive immune cells in the context of early metastasis are not well understood. Regulatory T cells and fibroblasts both have been shown to contribute to metastasis formation. We hypothesize that the IL-33/ST2 axis, through the interaction of IL-33 producing fibroblastic stromal cells and ST2-positive tissue resident regulatory T cells, plays an important role in early metastasis. The aim of this project is to functionally characterize the cellular and molecular mechanisms of the IL-33/ST2 axis important for early metastasis in order to develop novel interventional therapeutic strategies.
B07
The microenvironment at metastatic target sites determines the success of disseminating tumour cells to establish a metastatic colony. In metastasis of pancreatic ductal adenocarcinoma (PDAC) cancer associated fibroblasts (CAFs) are forming a very heterogeneous tumour microenvironment. This is reflected by activation of the EMT-inducing transcription factor ZEB1 in only a subset of CAFs. We hypothesize that ZEB1 is a crucial determinant of CAF subtype specification and function that drives disease progression. We will identify and analyse different CAF subpopulations and the role of ZEB1 in the process of colonization. We apply transplantation and genetic mouse tumour models, scRNA sequencing, ex vivo organ slice microscopy and human PDAC organoid-CAF co-cultures to address these hypothesis. With our findings on specific CAF subtypes, we hope to identify new therapeutic approaches targeting protumourigenic functions of the microenvironment during colonization.
B08
In colorectal cancer (CRC) distant metastases are the main cause of cancer-related death. The mutual interaction of disseminated cancer cells (DCCs) with stromal cells such as blood vessel endothelial cells is involved in the metastatic colonization. Recently, we have identified tumor microenvironment (TME)-dependent vascular plasticity in human CRC patients that actively supresses tumorigenesis by the release of the angiocrine protein SPARCL1 which exhibtis both, anti-angiogenic and anti-tumorigenic functions. In the proposed project B08, we will analyse whether SPARCL1 also inhibits the expansion of single DCCs and by this mechanism may actively counteract loco-regional and distant metastases. To these goals novel metastatic organoid-based CRC mouse models will be used. The long-term perspective is to establish SPARCL1 as a novel diagnostic marker to predict the risk of CRC metastasis and to elucidate the mechanisms of its anti-tumorigenic function to provide new targets for future therapeutic intervention.
B09
Melanoma spread to lymph nodes of the regional basin is a major predictor for melanoma outcome. Since lymph nodes are often resected, they offer the unique opportunity to catch human disseminated melanoma cells (DCCs) before and during metastatic colony formation and to study their interplay with local immune cells. Transcriptome analysis of DCCs identified activation of DCC-derived extracellular vesicle (EVs) production during colony formation, which was paralleled by CD8 T cell exhaustion, strongly suggesting a fundamental role of EVs in DCC-niche communication. We hypothesize that metastatic colony formation is driven by DCC-derived EVs, which modulate phenotype and function of surrounding immune cells and provide growth support to yet pre-colonizing DCC-clones and whose production is activated by oncogenic pathways. The goal of the project is to identify and mechanistically dissect the role of molecular constituents of DCC-derived EVs and oncogenic pathways involved therein.
B10
Several epidemiological studies suggest an association between obesity and both cancer incidence and mortality. In this context, we will compare the metastatic properties in lean and obese organisms using non-invasive imaging. In project B10, we will molecularly identify lean versus obese adipocyte/fat signaling from bone and liver that are required for the colonization of melanoma and breast cancer cells. In particular, we will delineate the importance of hypoxia, IL-6 signaling and presence of myeloid cells in lean versus obese adipose tissues. Finally, we will verify our results in patients with malignant melanoma and breast cancer. These experiments have a clear translational impact on the development of new therapeutic avenues for early detection and personalized treatment.
B11
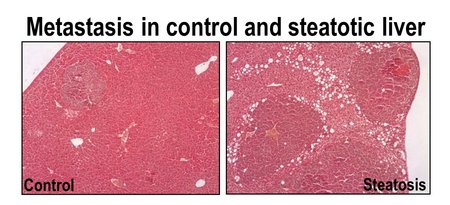
The liver is a highly metastasis-permissive organ. The molecular mechanisms that render the liver such hospitable to disseminated tumor cells are widely elusive. We found that the attractiveness of the liver for metastasizing cancer cells is further enhanced by hepatic steatosis. The general goal of this project is to identify the molecular mechanisms that promote the colonization of disseminated cancer cells into the steatotic hepatic environment and to validate their potential as novel therapeutic targets. We will also test whether identified factors affect colonization of non-steatotic liver tissue as well as other metastatic niches, where they may be less abundant but still effective.
B12
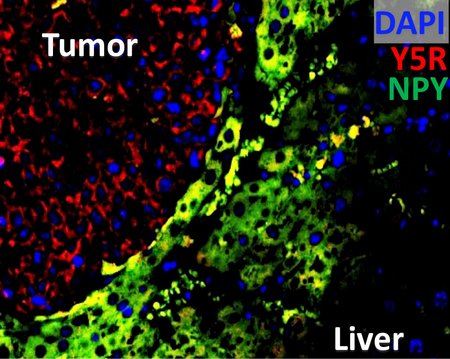
Liver organotropism is a hallmark of cancer. According to the "seed and soil" theory of metastasis, molecular cross-talk between cancer cells and non-tumorous parenchyma-derived cells underlies liver colonization. Our previous work and unpublished data reveal strong chemotactic effects of liver- and sympathetic-nerve-derived neuropeptide Y (Npy) on disseminated cancer cells that express Npy-receptors. The Npy/Npy-receptor cross-talk could drives homing, nerve infiltration and liver colonization by disseminated cancer cells. In this project, we aim at characterizing this molecular cross-talk between the liver metastatic niche and disseminated cancer cells applying in vitro and in vivo model systems as well as analyzing patient samples. Moreover, the Npy-dependent transcriptome in metastatic cells and its influence on EMT-induced gene signatures will be characterized. Furthermore, evaluation of potential therapeutic targetability of newly revealed pathways driving Npy-mediated metastasis will be addressed.
B13
To investigate early metastases and to test adjuvant therapies in presence of autologous immune cells novel preclinical immune competent in vitro models based on patient derived DCCs are needed. In project B13 we will therefore generate new LN-derived DCC organoid models from breast, colorectal and pancreatic cancer patients. To study colonization in a more complex and representative microenvironment we will use Precision Cut Tissue Slices from the main metastatic sites lung, liver and lymph nodes spiked with DCCs and immune cells from the same patients. To enable a prolonged cultivation of tissue slices and cocultures of DCCs in tissue preparations, we will develop new microfluidic systems. Thereby, we will establish a model of metastatic niches in vitro allowing to investigate mechanisms for colonization of DCCs in different organs, to unravel functions of different immune cells within this process and to test pharmaceutical interventions preventing colonization.
Z01
The core project Z01 will support the TRR 305 projects with the generation, analysis and handling of large datasets by data management, data sharing, bioinformatics, imaging and image analysis. We will allow effective data sharing across projects with an intuitive web-browser application and eventually make the joint data resources of the CRC publicly accessible with appropriate cross links between individual data sets. Furthermore, we will support all experimental CRC projects with expertise in the design of OMICS experiments, in the computational management and statistical analysis of the resulting data, and in the visualization and interpretation of the results. In addition, we will deliver advanced imaging infrastructures and tailored solutions for imaging of metastasis, as well as develop novel high-end image processing solutions for the TRR 305 members, including automated image analysis technologies.
Z02
The clinical-pathological core Z02 will establish a platform that serves as a unique resource to test, validate, and specify research questions addressed in the various subprojects and the TRR 305 in general. For this, Z02 will establish and provide two distinct resources: First, a sample repository addressing the needs of the TRR 305 projects and second, access and targeted exploitation of the prospective molecular registry study PRAEGNANT. The sample repository will comprise a clinical infrastructure for targeted post-mortem autopsy studies, logistics for high-quality targeted post-mortem sampling procedures and a platform to track prospectively and longitudinally disease courses to address questions of cellular and molecular cancer evolution. In addition, we will utilize the availability of samples of patients with metastasized breast cancer gathered during the prospective molecular registry study PRAEGNANT.